pygmt.grdtrack
- pygmt.grdtrack(grid, points=None, output_type='pandas', outfile=None, newcolname=None, region=None, verbose=False, incols=None, outcols=None, **kwargs)[source]
Sample one or more grids at specified locations.
Reads one or more grid files and a table (from file or an array input; but see
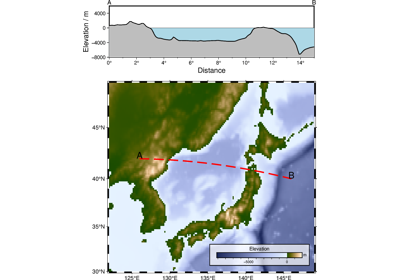
profilefor exception) with (x,y) [or (lon,lat)] positions in the first two columns (more columns may be present). It interpolates the grid(s) at the positions in the table and writes out the table with the interpolated values added as (one or more) new columns. Alternatively (crossprofile), the input is considered to be line-segments and we create orthogonal cross-profiles at each data point or with an equidistant separation and sample the grid(s) along these profiles. A bicubic [Default], bilinear, B-spline or nearest-neighbor interpolation is used, requiring boundary conditions at the limits of the region (seeinterpolation; Default uses “natural” conditions (second partial derivative normal to edge is zero) unless the grid is automatically recognized as periodic.)Full GMT docs at https://docs.generic-mapping-tools.org/6.6/grdtrack.html.
Aliases:
A = resample
C = crossprofile
D = dfile
E = profile
F = critical
N = no_skip
S = stack
T = radius
Z = z_only
a = aspatial
b = binary
d = nodata
e = find
f = coltypes
g = gap
h = header
j = distcalc
n = interpolation
s = skiprows
w = wrap
R = region
V = verbose
i = incols
o = outcols
- Parameters:
grid (
str|PathLike|DataArray) –Name of the input grid file or the grid loaded as a
xarray.DataArrayobject.For reading a specific grid file format or applying basic data operations, see https://docs.generic-mapping-tools.org/6.6/gmt.html#grd-inout-full for the available modifiers.
points (
str|PathLike|dict|ndarray|DataFrame|Dataset|GeoDataFrame|None, default:None) – Pass in either a file name to an ASCII data table, a 2-Dnumpy.ndarray, apandas.DataFrame, anxarray.Datasetmade up of 1-Dxarray.DataArraydata variables, or ageopandas.GeoDataFramecontaining the tabular data.output_type (
Literal['pandas','numpy','file'], default:'pandas') –Desired output type of the result data.
pandaswill return apandas.DataFrameobject.numpywill return anumpy.ndarrayobject.filewill save the result to the file specified by theoutfileparameter.
outfile (
str|PathLike|None, default:None) – File name for saving the result data. Required ifoutput_type="file". If specified,output_typewill be forced to be"file".newcolname (str) – Required if
pointsis apandas.DataFrame. The name for the new column in the trackpandas.DataFrametable where the sampled values will be placed.resample (str) – f|p|m|r|R[+l] For track resampling (if
crossprofileorprofileare set) we can select how this is to be performed. Append f to keep original points, but add intermediate points if needed [Default], m as f, but first follow meridian (along y) then parallel (along x), p as f, but first follow parallel (along y) then meridian (along x), r to resample at equidistant locations; input points are not necessarily included in the output, and R as r, but adjust given spacing to fit the track length exactly. Finally, append +l if geographic distances should be measured along rhumb lines (loxodromes) instead of great circles. Ignored unlesscrossprofileis used.crossprofile (str) – length/ds[/spacing][+a|+v][l|r]. Use input line segments to create an equidistant and (optionally) equally-spaced set of crossing profiles along which we sample the grid(s) [Default simply samples the grid(s) at the input locations]. Specify two length scales that control how the sampling is done: length sets the full length of each cross-profile, while ds is the sampling spacing along each cross-profile. Optionally, append /spacing for an equidistant spacing between cross-profiles [Default erects cross-profiles at the input coordinates]; see
resamplefor how resampling the input track is controlled. By default, all cross-profiles have the same direction (left to right as we look in the direction of the input line segment). Append +a to alternate the direction of cross-profiles, or v to enforce either a “west-to-east” or “south-to-north” view. By default the entire profiles are output. Choose to only output the left or right halves of the profiles by appending +l or +r, respectively. Append suitable units to length; it sets the unit used for ds [and spacing] (See Units). The default unit for geographic grids is meters while Cartesian grids implies the user unit. The output columns will be lon, lat, dist, azimuth, z1, z2, …, zn (The zi are the sampled values for each of the n grids).dfile (str) – In concert with
crossprofilewe can save the (possibly resampled) original lines to dfile [Default only saves the cross-profiles]. The columns will be lon, lat, dist, azimuth, z1, z2, … (sampled value for each grid).profile (str) – line[,line,…][+aaz][+c][+d][+g][+iinc][+llength][+nnp][+oaz][+rradius]. Instead of reading input track coordinates, specify profiles via coordinates and modifiers. The format of each line is start/stop, where start or stop are either lon/lat (x/y for Cartesian data) or a 2-character XY key that uses the text-style justification format to specify a point on the map as [LCR][BMT]. Each line will be a separate segment unless +c is used which will connect segments with shared joints into a single segment. In addition to line coordinates, you can use Z-, Z+ to mean the global minimum and maximum locations in the grid (only available if a single grid is given via outfile). You may append +iinc to set the sampling interval; if not given then we default to half the minimum grid interval. For a line along parallels or meridians you can add +g to report degrees of longitude or latitude instead of great circle distances starting at zero. Instead of two coordinates you can specify an origin and one of +a, +o, or +r. The +a sets the azimuth of a profile of given length starting at the given origin, while +o centers the profile on the origin; both require +l. For circular sampling specify +r to define a circle of given radius centered on the origin; this option requires either +n or +i. The +nnp modifier sets the desired number of points, while +llength gives the total length of the profile. Append +d to output the along-track distances after the coordinates. Note: No track file will be read. Also note that only one distance unit can be chosen. Giving different units will result in an error. If no units are specified we default to great circle distances in km (if geographic). If working with geographic data you can use
distcalcto control distance calculation mode [Default is Great Circle]. Note: Ifcrossprofileis set and spacing is given then that sampling scheme overrules any modifier set inprofile.critical (str) – [+b][+n][+r][+zz0]. Find critical points along each cross-profile as a function of along-track distance. Requires
crossprofileand a single input grid (z). We examine each cross-profile generated and report (dist, lonc, latc, distc, azimuthc, zc) at the center peak of maximum z value, (lonl, latl, distl) and (lonr, latr, distr) at the first and last non-NaN point whose z-value exceeds z0, respectively, and the width based on the two extreme points found. Here, dist is the distance along the original inputpointsand the other 12 output columns are a function of that distance. When searching for the center peak and the extreme first and last values that exceed the threshold we assume the profile is positive up. If we instead are looking for a trough then you must use +n to temporarily flip the profile to positive. The threshold z0 value is always given as >= 0; use +z to change it [Default is 0]. Alternatively, use +b to determine the balance point and standard deviation of the profile; this is the weighted mean and weighted standard deviation of the distances, with z acting as the weight. Finally, use +r to obtain the weighted rms about the cross-track center (distc == 0). Note: We round the exact results to the nearest distance nodes along the cross-profiles. We write 13 output columns per track: dist, lonc, latc, distc, azimuthc, zc, lonl, latl, distl, lonr, latr, distr, width.region (str or list) – xmin/xmax/ymin/ymax[+r][+uunit]. Specify the region of interest.
no_skip (bool) – Do not skip points that fall outside the domain of the grid(s) [Default only output points within the grid domain].
method/modifiers. In conjunction with
crossprofile, compute a single stacked profile from all profiles across each segment. Choose how stacking should be computed [Default method is a]:a = mean (average)
m = median
p = mode (maximum likelihood)
l = lower
L = lower but only consider positive values
u = upper
U = upper but only consider negative values.
The modifiers control the output; choose one or more among these choices:
+a : Append stacked values to all cross-profiles.
+d : Append stack deviations to all cross-profiles.
+r : Append data residuals (data - stack) to all cross-profiles.
+s[file] : Save stacked profile to file [Default file name is grdtrack_stacked_profile.txt].
+cfact : Compute envelope on stacked profile as ±fact *deviation [Default fact value is 2].
Here are some notes:
Deviations depend on method and are st.dev (a), L1 scale, i.e., 1.4826 * median absolute deviation (MAD) (for m and p), or half-range (upper-lower)/2.
The stacked profile file contains a leading column plus groups of 4-6 columns, with one group for each sampled grid. The leading column holds cross distance, while the first four columns in a group hold stacked value, deviation, min value, and max value, respectively. If method is one of a|m|p then we also write the lower and upper confidence bounds (see +c). When one or more of +a, +d, and +r are used then we also append the stacking results to the end of each row, for all cross-profiles. The order is always stacked value (+a), followed by deviations (+d) and finally residuals (+r). When more than one grid is sampled this sequence of 1-3 columns is repeated for each grid.
radius (bool, float, or str) – [radius][+e|p]. To be used with normal grid sampling, and limited to a single, non-IMG grid. If the nearest node to the input point is NaN, search outwards until we find the nearest non-NaN node and report that value instead. Optionally specify a search radius which limits the consideration to points within this distance from the input point. To report the location of the nearest node and its distance from the input point, append +e. The default unit for geographic grid distances is spherical degrees. Use radius to change the unit and give radius = 0 if you do not want to limit the radius search. To instead replace the input point with the coordinates of the nearest node, append +p.
verbose (bool or str) – Select verbosity level [Full usage].
z_only (bool) – Only write out the sampled z-values [Default writes all columns].
aspatial (bool or str) – [col=]name[,…]. Control how aspatial data are handled during input and output. Full documentation is at https://docs.generic-mapping-tools.org/6.6/gmt.html#aspatial-full.
i|o[ncols][type][w][+l|b]. Select native binary input (using
binary="i") or output (usingbinary="o"), where ncols is the number of data columns of type, which must be one of:c: int8_t (1-byte signed char)
u: uint8_t (1-byte unsigned char)
h: int16_t (2-byte signed int)
H: uint16_t (2-byte unsigned int)
i: int32_t (4-byte signed int)
I: uint32_t (4-byte unsigned int)
l: int64_t (8-byte signed int)
L: uint64_t (8-byte unsigned int)
f: 4-byte single-precision float
d: 8-byte double-precision float
x: use to skip ncols anywhere in the record
For records with mixed types, append additional comma-separated combinations of ncols type (no space). The following modifiers are supported:
w after any item to force byte-swapping.
+l|b to indicate that the entire data file should be read as little- or big-endian, respectively.
Full documentation is at https://docs.generic-mapping-tools.org/6.6/gmt.html#bi-full.
nodata (str) – i|onodata. Substitute specific values with NaN (for tabular data). For example,
nodata="-9999"will replace all values equal to -9999 with NaN during input and all NaN values with -9999 during output. Prepend i to the nodata value for input columns only. Prepend o to the nodata value for output columns only.find (str) – [~]“pattern” | [~]/regexp/[i]. Only pass records that match the given pattern or regular expressions [Default processes all records]. Prepend ~ to the pattern or regexp to instead only pass data expressions that do not match the pattern. Append i for case insensitive matching. This does not apply to headers or segment headers.
coltypes (str) – [i|o]colinfo. Specify data types of input and/or output columns (time or geographical data). Full documentation is at https://docs.generic-mapping-tools.org/6.6/gmt.html#f-full.
x|y|z|d|X|Y|Dgap[u][+a][+ccol][+n|p]. Examine the spacing between consecutive data points in order to impose breaks in the line. To specify multiple criteria, provide a list with each item containing a string describing one set of criteria.
x|X: define a gap when there is a large enough change in the x coordinates (uppercase to use projected coordinates).
y|Y: define a gap when there is a large enough change in the y coordinates (uppercase to use projected coordinates).
d|D: define a gap when there is a large enough distance between coordinates (uppercase to use projected coordinates).
z: define a gap when there is a large enough change in the z data. Use +ccol to change the z data column [Default col is 2 (i.e., 3rd column)].
A unit u may be appended to the specified gap:
For geographic data (x|y|d), the unit may be arc- d(egrees), m(inutes), and s(econds), or (m)e(ters), f(eet), k(ilometers), M(iles), or n(autical miles) [Default is (m)e(ters)].
For projected data (X|Y|D), the unit may be i(nches), c(entimeters), or p(oints).
Append modifier +a to specify that all the criteria must be met [default imposes breaks if any one criterion is met].
One of the following modifiers can be appended:
+n: specify that the previous value minus the current column value must exceed gap for a break to be imposed.
+p: specify that the current value minus the previous value must exceed gap for a break to be imposed.
header (str) –
[i|o][n][+c][+d][+msegheader][+rremark][+ttitle]. Specify that input and/or output file(s) have n header records [Default is 0]. Prepend i if only the primary input should have header records. Prepend o to control the writing of header records, with the following modifiers supported:
+d to remove existing header records.
+c to add a header comment with column names to the output [Default is no column names].
+m to add a segment header segheader to the output after the header block [Default is no segment header].
+r to add a remark comment to the output [Default is no comment]. The remark string may contain \n to indicate line-breaks.
+t to add a title comment to the output [Default is no title]. The title string may contain \n to indicate line-breaks.
Blank lines and lines starting with # are always skipped.
incols (
int|str|Sequence[int|str] |None, default:None) –Specify data columns for primary input in arbitrary order. Columns can be repeated and columns not listed will be skipped [Default reads all columns in order, starting with the first (i.e., column 0)].
For a sequence: specify individual columns in input order (e.g.,
incols=(1, 0)for the 2nd column followed by the 1st column).For a string: specify individual columns or column ranges in the format start[:inc]:stop, where inc defaults to 1 if not specified, with columns and/or column ranges separated by commas (e.g.,
incols="0:2,4+l"to input the first three columns followed by the log10-transformed 5th column). To read from a given column until the end of the record, leave off stop when specifying the column range. To read trailing text, add the column t. Append the word number to t to ingest only a single word from the trailing text. Instead of specifying columns, useincols="n"to simply read numerical input and skip trailing text. Optionally, append one of the following modifiers to any column or column range to transform the input columns:+l to take the log10 of the input values.
+d to divide the input values by the factor divisor [Default is 1].
+s to multiple the input values by the factor scale [Default is 1].
+o to add the given offset to the input values [Default is 0].
distcalc (str) – Determine how spherical distances are calculated [Full usage].
interpolation (str) –
[b|c|l|n][+a][+bBC][+c][+tthreshold]. Select interpolation mode for grids. You can select the type of spline used:
b for B-spline
c for bicubic [Default]
l for bilinear
n for nearest-neighbor
outcols (
int|str|Sequence[int|str] |None, default:None) –cols[,…][,t[word]].
Specify data columns for primary output in arbitrary order. Columns can be repeated and columns not listed will be skipped [Default writes all columns in order, starting with the first (i.e., column 0)].
For a sequence: specify individual columns in output order (e.g.,
outcols=(1, 0)for the 2nd column followed by the 1st column).For a string: specify individual columns or column ranges in the format start[:inc]:stop, where inc defaults to 1 if not specified, with columns and/or column ranges separated by commas (e.g.,
outcols="0:2,4"to output the first three columns followed by the 5th column). To write from a given column until the end of the record, leave off stop when specifying the column range. To write trailing text, add the column t. Append the word number to t to write only a single word from the trailing text. Instead of specifying columns, useoutcols="n"to simply read numerical input and skip trailing text. Note: Ifincolsis also used then the columns given tooutcolscorrespond to the order after theincolsselection has taken place.Optionally, append one of the following modifiers to any column or column range to transform the output columns:
+l to take the log10 of the input values.
+d to divide the output values by the factor divisor [Default is 1].
+s to multiply the output values by the factor scale [Default is 1].
+o to add the given offset to the output values [Default is 0].
[cols][+a][+r]. Suppress output for records whose z-value equals NaN [Default outputs all records]. Optionally, supply a comma-separated list of all columns or column ranges to consider for this NaN test [Default only considers the third data column (i.e., cols = 2)]. Column ranges must be given in the format start[:inc]:stop, where inc defaults to 1 if not specified. The following modifiers are supported:
+r to reverse the suppression, i.e., only output the records whose z-value equals NaN.
+a to suppress the output of the record if just one or more of the columns equal NaN [Default skips record only if values in all specified cols equal NaN].
wrap (str) –
y|a|w|d|h|m|s|cperiod[/phase][+ccol]. Convert the input x-coordinate to a cyclical coordinate, or a different column if selected via +ccol. The following cyclical coordinate transformations are supported:
y: yearly cycle (normalized)
a: annual cycle (monthly)
w: weekly cycle (day)
d: daily cycle (hour)
h: hourly cycle (minute)
m: minute cycle (second)
s: second cycle (second)
c: custom cycle (normalized)
Full documentation is at https://docs.generic-mapping-tools.org/6.6/gmt.html#w-full.
- Return type:
- Returns:
ret – Return type depends on
outfileandoutput_type:Noneifoutfileis set (output will be stored in file set byoutfile)pandas.DataFrameornumpy.ndarrayifoutfileis not set (depends onoutput_type)
Example
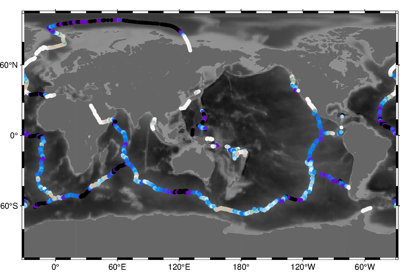
>>> import pygmt >>> # Load a grid of @earth_relief_30m data, with a longitude range of >>> # -118° E to -107° E, and a latitude range of -49° N to -42° N >>> grid = pygmt.datasets.load_earth_relief( ... resolution="30m", region=[-118, -107, -49, -42] ... ) >>> # Load a pandas dataframe with ocean ridge points >>> points = pygmt.datasets.load_sample_data(name="ocean_ridge_points") >>> # Create a pandas dataframe from an input grid and set of points >>> # The output dataframe adds a column named "bathymetry" >>> output_dataframe = pygmt.grdtrack( ... points=points, grid=grid, newcolname="bathymetry" ... )